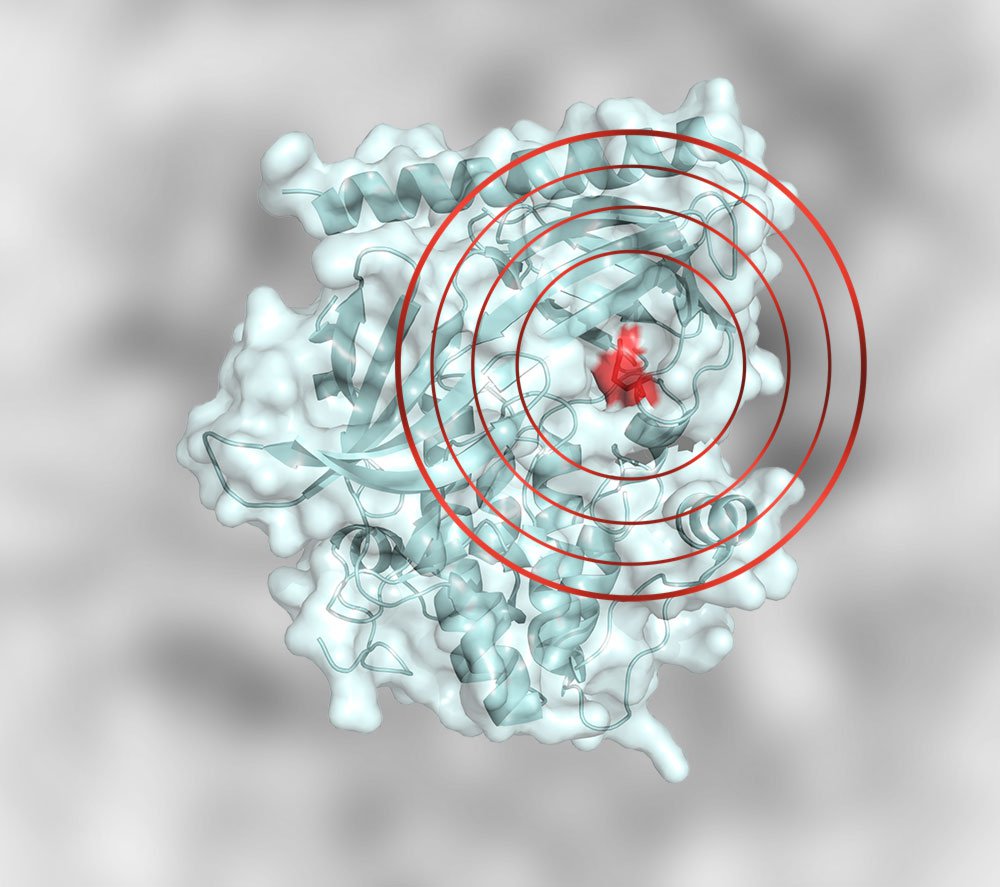
WDR76 is a RAS binding protein that functions as a tumor suppressor via RAS degradation | Nature Communications

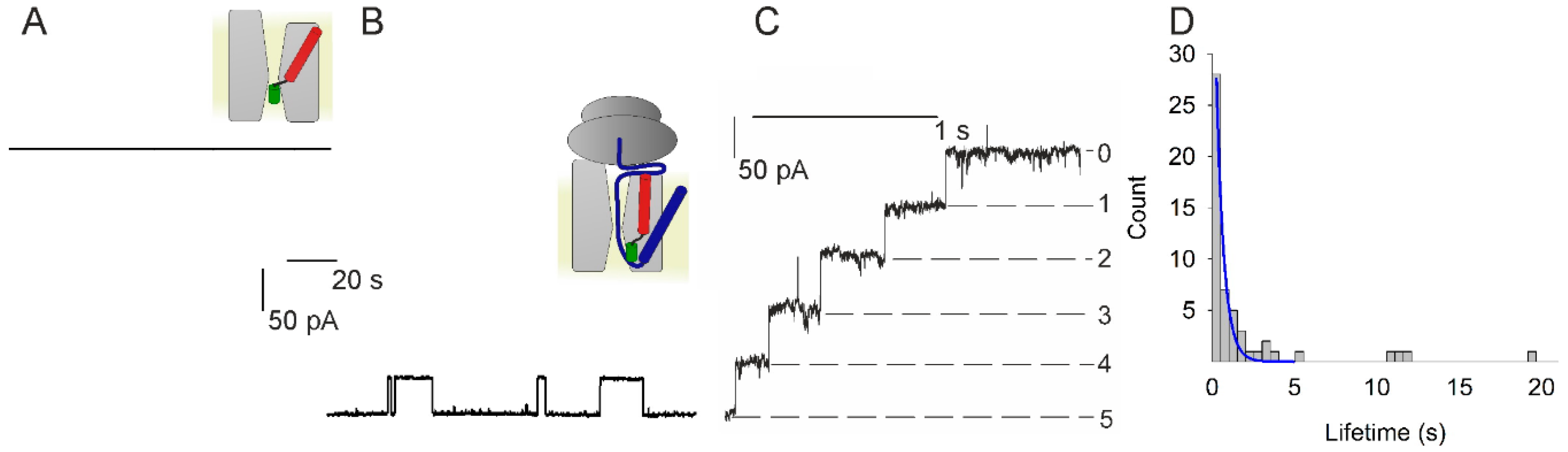
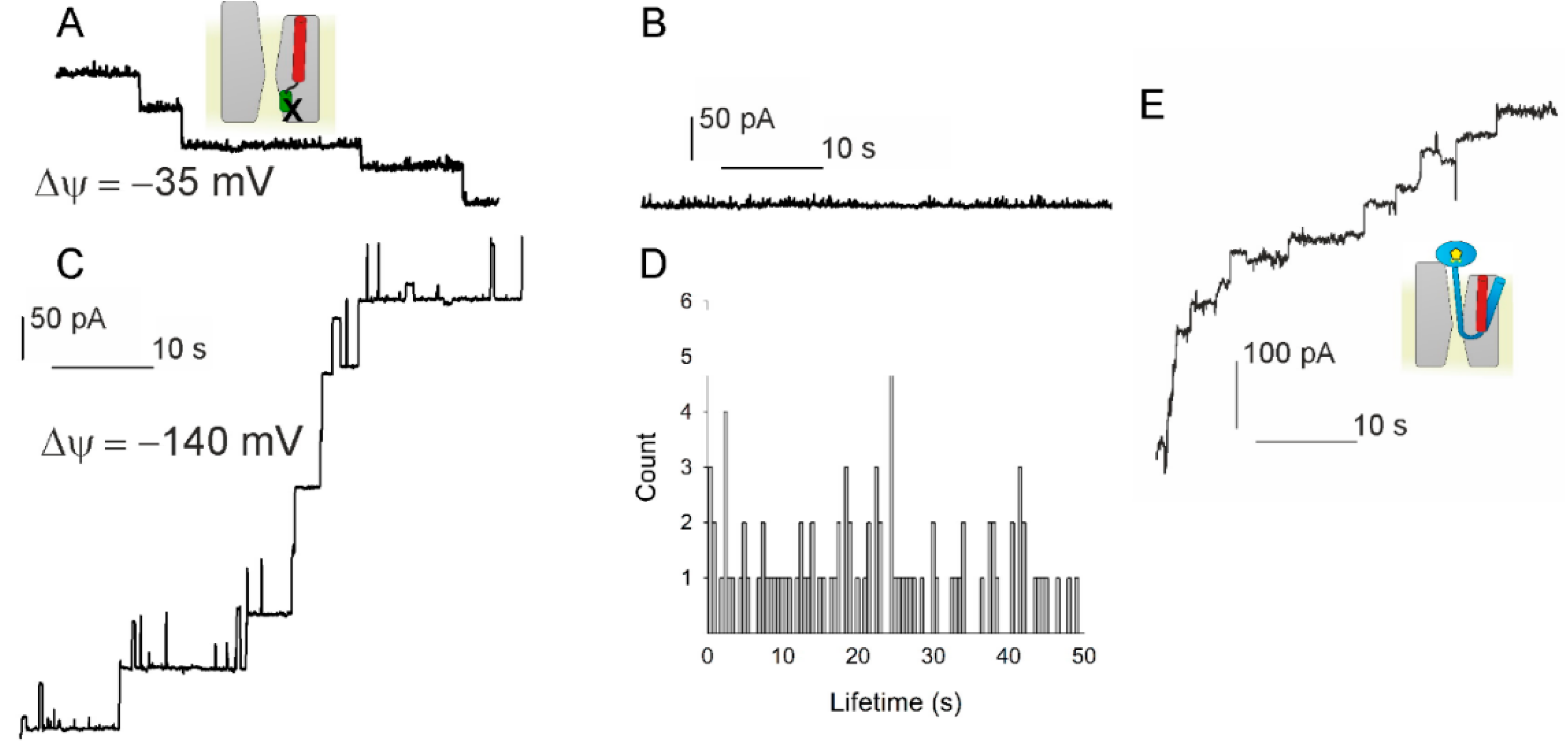
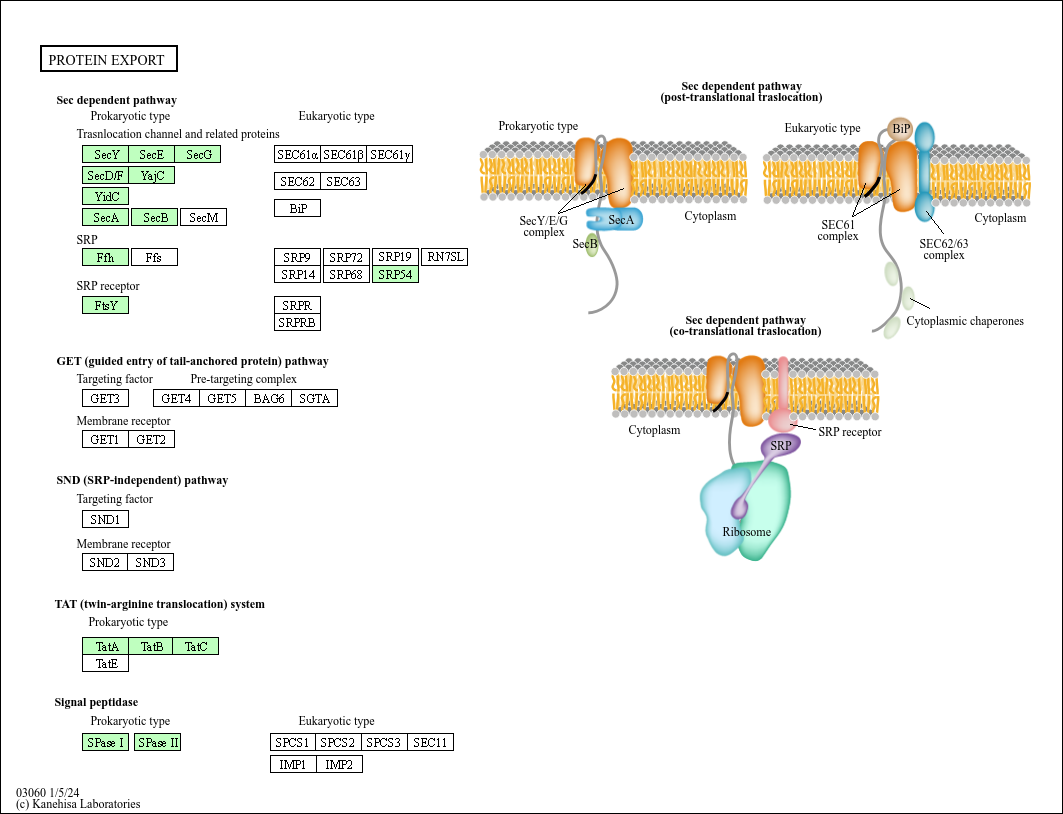
Cotranslational protein targeting to the membrane: Nascent-chain transfer in a quaternary complex formed at the translocon | Scientific Reports

FKBP25/FKBP3 Protein Overview: Sequence, Structure, Function and Protein Interaction | Sino Biological

RNA-binding proteins and heat-shock protein 90 are constituents of the cytoplasmic capping enzyme interactome - Journal of Biological Chemistry

Adeno-Associated Virus Type 2-Mediated Gene Transfer: Role of Cellular FKBP52 Protein in Transgene Expression | Journal of Virology

Protein disulfide isomerase externalization in endothelial cells follows classical and unconventional routes - ScienceDirect
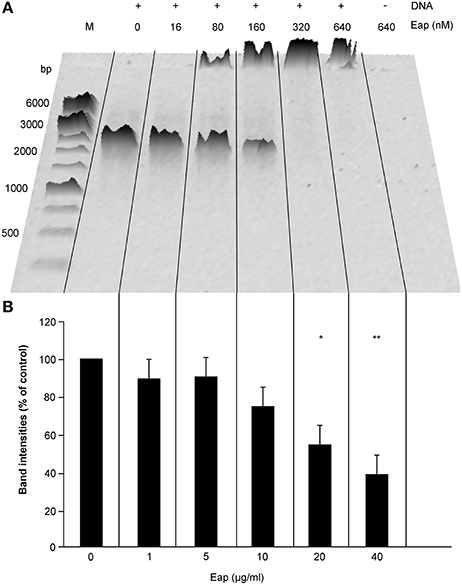
Frontiers | The Staphylococcus aureus Extracellular Adherence Protein Eap Is a DNA Binding Protein Capable of Blocking Neutrophil Extracellular Trap Formation | Cellular and Infection Microbiology
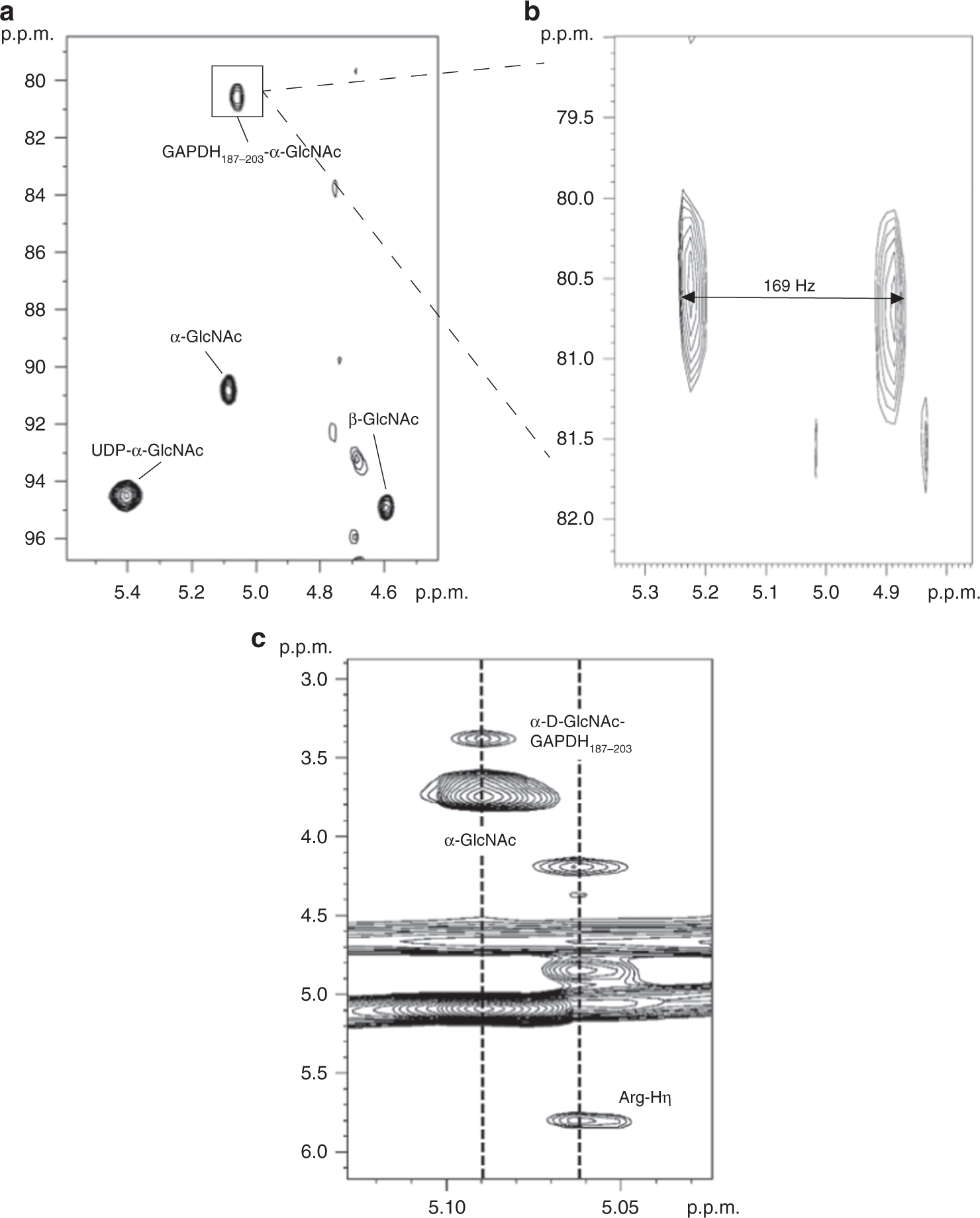
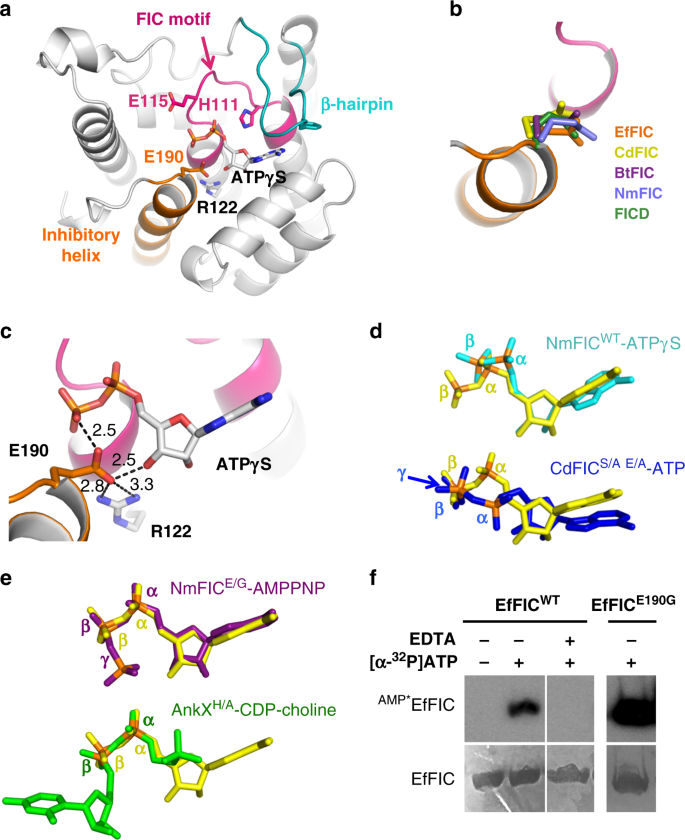
Structural basis for arginine glycosylation of host substrates by bacterial effector proteins | Nature Communications

Structure, gene expression, and putative functions of crustacean heat shock proteins in innate immunity - ScienceDirect
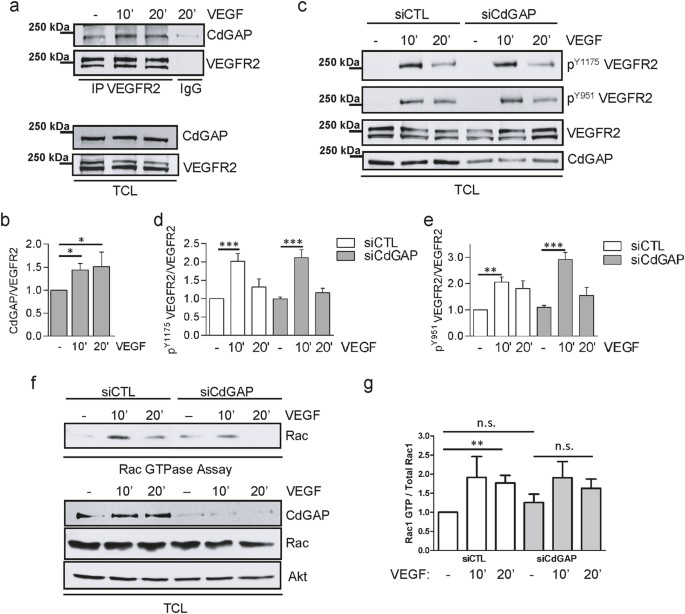
CdGAP/ARHGAP31, a Cdc42/Rac1 GTPase regulator, is critical for vascular development and VEGF-mediated angiogenesis | Scientific Reports
Poster session abstracts – topic of research paper in Biological sciences. Download scholarly article PDF and read for free on CyberLeninka open science hub.

Role of Nucleotide Binding and GTPase Domain Dimerization in Dynamin-like Myxovirus Resistance Protein A for GTPase Activation and Antiviral Activity* - Journal of Biological Chemistry